Wu and Carson [1] aimed at making the SRTM basis function approach even more robust and called it Simplified Reference Tissue Model 2 (SRTM2). They noted that with SRTM k2' is calculated with each pixel TAC, although the same reference TAC is used for all pixels. Therefore they implemented a two-step approach:
1.Calculate k2' using SRTM in all pixels.
2.Fix k2' : Average k2' in all brain pixels outside the reference region. Use this fixed value for the pixelwise SRTM calculations, reducing the number of fitted parameters from 3 to 2.
The operational equation of the SRTM was re-written to allow for fixing of k2'. This is relevant for parametric mapping because the model in each pixel TAC uses the same reference TAC and therefore should employ the same k2'. Defining the ratio of tracer delivery R1 as K1/K1' and the binding potential BPND as k3/k4, the following operational equation can be derived for the measured TAC in a receptor-rich region:
![]()
The three unknowns R1, k2' and k2a in this equation can be fitted using nonlinear regression techniques. The binding potential can then be calculated as
![]()
The current PXMOD implementation BPnd (Wu SRTM2 Ref) supports four methods for specifying k2' as described in section Specification of k2'.
Acquisition and Data Requirements
Image Data |
A dynamic PET data set imaging a receptor tracer which behaves kinetically similar to a 1-tissue compartment model. |
Target tissue |
Optional: TAC from a receptor-rich region (such as basal ganglia for D2 receptors). For visualization of model fitting and optionally for fitting k2' |
Reference tissue |
Mandatory: TAC from a receptor-devoid reference region (such as cerebellum or frontal cortex for D2 receptors). Note: specification of an appropriate reference TAC is crucial for the result! |
VOI |
Optional: VOI definition excluding the reference tissue which can be used for getting an estimate of k2'. |
Model Preprocessing
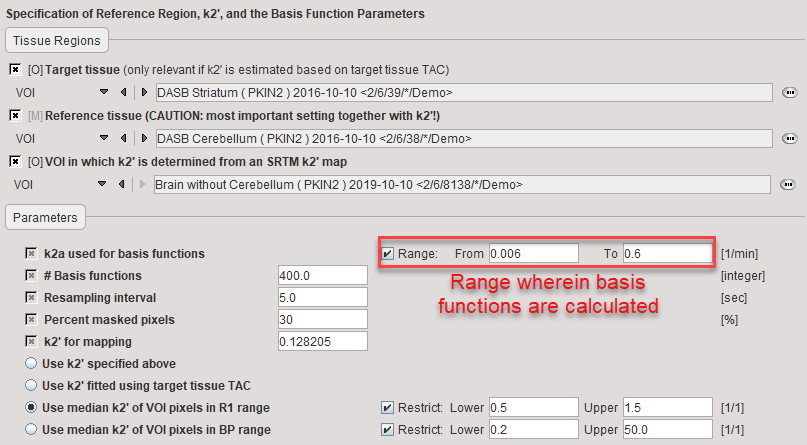
In the lower part the following parameters are configured:
k2a used for basis functions |
Enter the minimal value of k2a (slowest decay of exponential) and the maximal value of k2a (fastest decay of exponential) after checking the Range box. |
# Basis functions |
Number of basis functions between the minima and maximal k2a. Note that increments are taken at logarithmic steps. This number is directly proportional to processing time. |
Resampling interval |
Specifies the interval of curve resampling which is required for performing the operation of exponential convolution. Resampling interval should be equal or smaller than the shortest frame duration. |
Percent masked pixels |
Exclude the specified percentage of pixels based on histogram analysis of integrated signal energy. Not applied in the presence of a defined mask. |
k2' for mapping |
k2 of the reference tissue. It can be specified in four different ways, as explained in section Specification of k2'. |
The result of a model fit during Model Preprocessing is shown in the Model Results panel for inspection. If no Target tissue is specified, the panel remains empty.
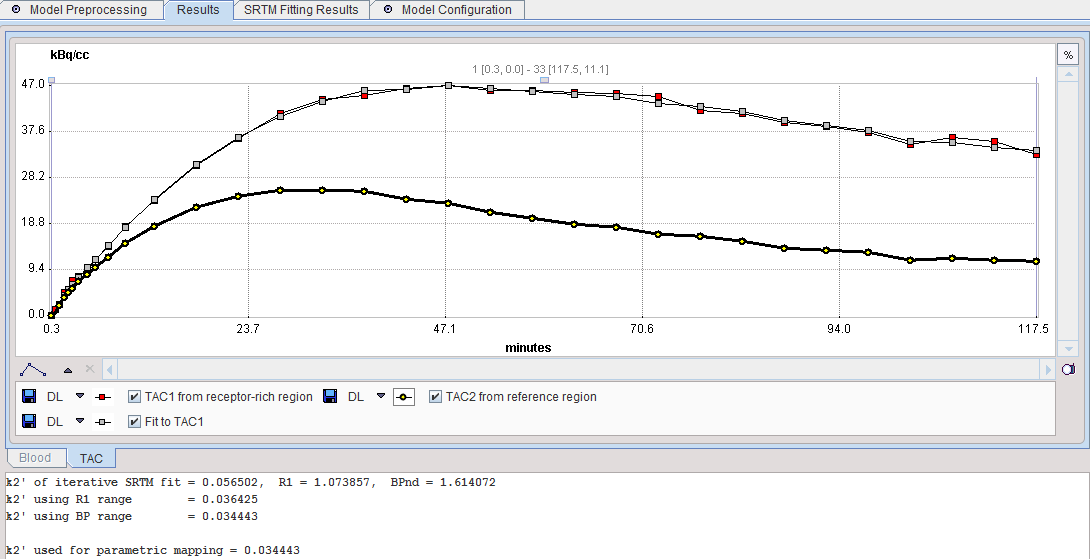
Note the lower text section which lists the k2' results of all three procedures as well as the k2' configured for the pixelwise analysis.
The model features an additional panel SRTM Fitting Results which allows inspecting the SRTM parametric maps with in the VOI. The parameter can be switched in the upper right. Due to outlier results it may be required to manually enter a physiologic value as the upper image threshold as illustrated below.
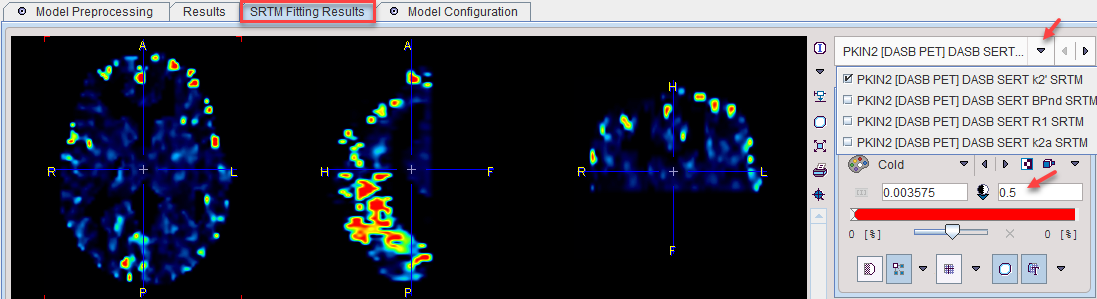
Model Configuration
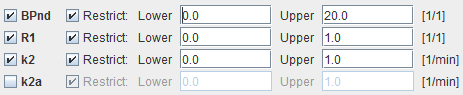
BPnd |
Estimated binding potential (BPnd= k3/k4 according to the underlying model). |
k2 |
Estimated efflux rate constant k2 . |
R1 |
Ratio of tracer delivery in each pixel relative to the reference tissue (R1=K1/K1'). Therefore the map often has a similar appearance to a perfusion image. |
k2a |
k2a value which provides the best least squares fit. |
Note: The k2a parametric map should be checked in the initial setup of a processing protocol. The estimated k2a values should not be truncated by too narrow k2a min and k2a max values.
Reference
1.Wu Y, Carson RE: Noise reduction in the simplified reference tissue model for neuroreceptor functional imaging. J Cereb Blood Flow Metab 2002, 22(12):1440-1452. DOI